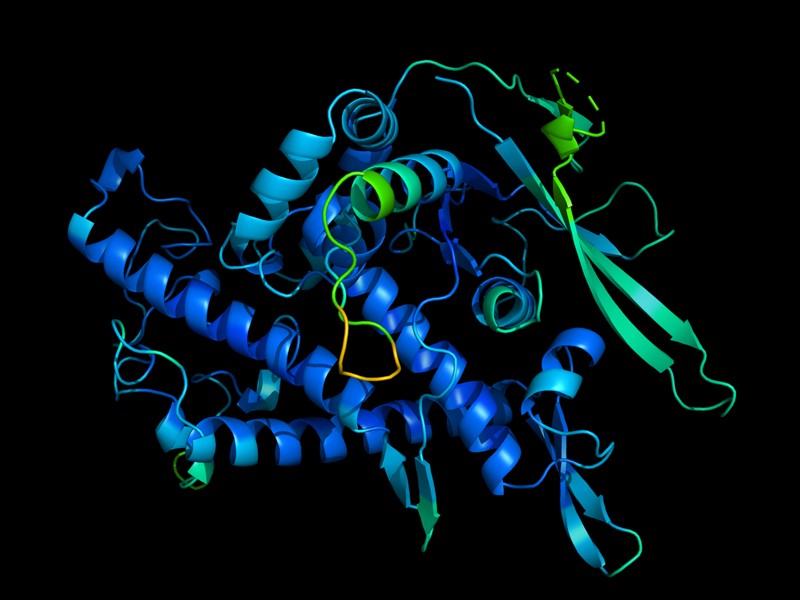
Proteins are behind every biological reaction happening within the human body, helping us ensure our immune system’s health. Out of many properties, one that allows them functioning well and fulfill their various tasks is their shape. They have a raveled three-dimensional shape that acquires via ‘protein folding’ – a process in which sequences of amino acids, the building blocks of protein, interact with one another to ‘fold’ into proteins. The 3-D structure of a protein determines its function. The underlying cause of many diseases can be that the protein is misfolded.
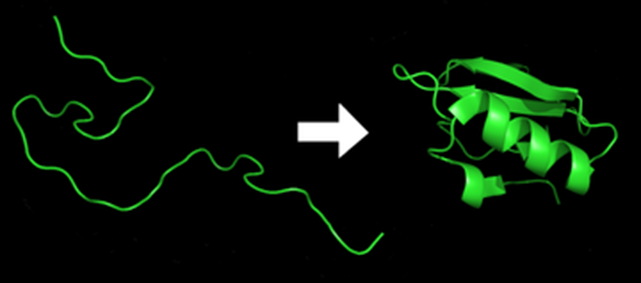
THE PROTEIN FOLDING PROBLEM
In more than 200 million proteins we are aware of, we know a few structures so far. There are various methods to predict the structures of proteins ranging from x-ray crystallography to nuclear magnetic resonance. But progress has been slow and heavy on the budget. X-ray crystallography costs around 120,000 dollars and can take up to a year to give a result. We can better understand the difficulty in predicting a protein structure through Levinthal’s paradox. In 1969, Cyrus Levinthal – an American molecular biologist – claimed that, according to the calculations, the protein folding process should take place over a period longer than the age of our universe. Yet proteins usually fold within a millisecond.
STORY OF CASP
The fifty-year-old ‘protein folding problem’ seemed to be far away from advancing until now. Out of nowhere, Google’s company DeepMind has come up with a way to demystify the different structures of proteins, understanding their function in the process! It can justly be considered one of the significant breakthroughs in structural biology as well as artificial intelligence.
The story begins in 1994 when the first edition of Critical Assessment of protein Structure Prediction, or CASP (I recommend using this), was held. Since then, the community-driven experiment has happened every two years. Every edition, teams participating are given a hundred proteins that have already mapped using one of the methods mentioned above. The groups, who don’t have access to the mapped structure, submit their predicted arrangement to be reviewed by the global distance test, or GDT for short. They are compared to the pre-existing structure to find measure homogeneity.
EMERGENCE OF ALPHAFOLD
In 2018, DeepMind participated in CASP13, where they triumphed with their algorithm – Alphafold. They surpassed expectations with the most accurate predictions till then, leaving the competition behind. Researchers were impressed by the results of Alphafold and deemed it to be a great success. But for DeepMind, this was just the beginning. 2020 saw CASP14 being held virtually due to the ongoing pandemic. DeepMind submitted their structures predicted by Alphafold 2, a neural network system trained by being fed a large amount of data on protein structures and their constituent amino acid sequences.
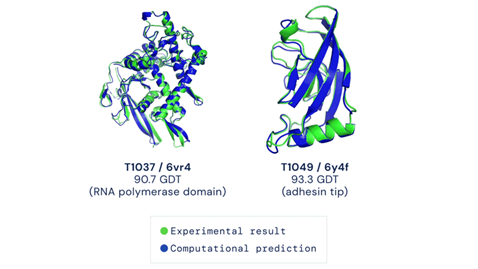
The results that came out were unprecedented. A score above 90 on the GDT is considered accurate and matches the best experimental methods’ standard. In other words, a score above 90 means that the problem is solved. Alphafold 2 scored between 87 to 92.4! As can be inferred, this is a big step in solving the ‘protein-fold problem’ taken via the unlikeliest of routes – Artificial Intelligence.
WHY IS THIS SO IMPORTANT?
Alphafold 2 could have far-reaching implications spanning across many fields. Molecular biologists will now be able to advance onto more complex and specific questions with access to reliable protein structure predictions. With a better understanding of the way proteins fold, researchers will better understand drug discovery. They could design specific medicines for diseases with newfound knowledge about protein structures and their functions. This year, Alphafold successfully predicted several protein structures of the SARS-CoV-2, the virus behind COVID-19, previously unknown.
DeepMind is looking forward to collaborating with biologists and researchers to realize the impact of Alphafold 2 further. They also aim to make the system more accessible for research labs. Needless to say, DeepMind has advanced not only the pursuit of molecular biology but also the pursuit of computation and AI. The success story of DeepMind is a welcomed one and holds even more significance due to the challenging times it has come in.
Bibliography
- https://deepmind.com/blog/article/AlphaFold-Using-AI-for-scientific-discovery
- https://deepmind.com/blog/article/alphafold-a-solution-to-a-50-year-old-grand-challenge-in-biology
Also Read: Philanthropic contribution in the healthcare system of Pakistan

Shoaib Shahid is a high school student doing his A-levels from Beaconhouse Margalla Islamabad. He is a great admirer of Physics and aspires to become a theoretical physicist himself. He started writing narratives in his childhood and has since ventured into non-fiction.